- Blog
- At home heart monitor
- 2020 leap year
- Sylo coin market cap
- Pinball wizard position
- Media room carpet ideas
- Proud boys flag
- Cellprofiler pipelines
- A world for two with you in the middle
- Battleship online vs computer
- Tyrion lannister quotes s07e03
- Star wars silver screen edition 1-6
- Synth 76477 ipad
- Ghost browser multiple user
- Discord logo maker online
- Does instagram notify of screenshots

- #CELLPROFILER PIPELINES FULL#
- #CELLPROFILER PIPELINES SOFTWARE#
- #CELLPROFILER PIPELINES FREE#
- #CELLPROFILER PIPELINES WINDOWS#
In addition, we provide CellProfiler pipelines for endogenous SQSTM1/p62.
#CELLPROFILER PIPELINES SOFTWARE#
(see file structure above)ĥ) Collect measurements using CellProfiler: bash scripts/run_cellprofiler.sh projects/20191119_example batch_files/20191119_example_batch_.h5īash scripts/check_run_cellprofiler.sh projects/20191119_example batch_files/20191119_example_batch_.h5 Ħ) Aggregate measurement data: Rscript scripts/aggregate_cellprofiler_results.R 20191119_example_data_ 20191119_example_metadata_.csv Īt this point, a summarized. The CellProfiler protocol is provided as a ready-to-use software pipeline.
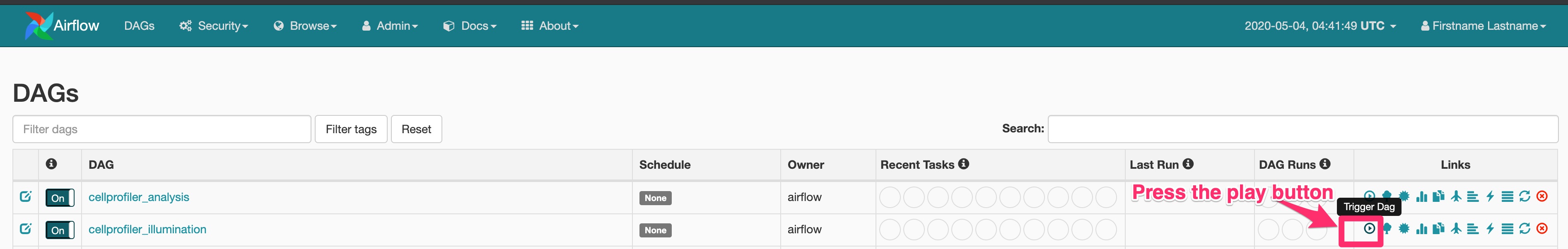
#CELLPROFILER PIPELINES FULL#
To execute the CP pipeline on QUEST: ¶ Full instructions here: ¶ġ) Navigate to the CP Directory: cd CellProfilerĢ) Generate metadata CP file: Rscript scripts/generate_metadata.R 20191119_exampleģ) Download metadata file and create CellProfiler pipeline locally.Ĥ) Upload properly named batch file and CellProfiler pipeline to appropriate QUEST directories. Once segmented, over 100 features of microtubule and chromatin structures were determined by CellProfiler modules, which resulted in an extensive dataset. image segmentation by combining Ilastik with CellProfiler was based on the analysis pipeline as described by the. Singularity pull docker://cellprofiler/cellprofiler:4.0.3 Singularity pull docker://cellprofiler/cellprofiler:3.1.9 To recreate CellProfiler (CP) on QUEST: ¶ġ) Navigate to the directory where you wish to clone CP: ssh -X ģ) Download CP Docker images and convert to Singularity images within CP directory: cd CellProfiler Thanks to David Stirling, Alice Lucas, Allen Goodman, Vito Zanotelli, Lijun Wang, Erin Weisbart, and Martin Dahlo for contributing to CellProfiler and/or CellProfiler-core since our 4.Andersen Lab Image Analysis Pipeline ¶ Implemented using CellProfiler ¶ CellProfiler Directory Structure (With Example Files) ¶ CellProfiler This can be done even if a pipeline is already loaded into that section. We’ve also moved to BioFormats 6.6.0 and made a number of small bug fixes, including to the display functions for several 3D modules, to modules that were using Minimum-Cross-Entropy thresholding (where occasionally images would cause the program to hang indefinitely), to measurements on Secondary objects when they were filtered for touching the edge but their parent objects were not, and to the MeasureImageQuality and Align modules. Previously created pipelines may be opened by dragging and dropping them into the section of CellProfiler indicated by the red arrow below. limited to) those produced by CellProfiler pipelines (Jones et al., 2008).
#CELLPROFILER PIPELINES WINDOWS#
We’ve made it easier to co-install multiple versions of CellProfiler on the same Windows machine by changing our Java packaging. CellProfiler Analyst is a free, open-source software package designed for the.Pipeline() file /notebooks/CellProfiler/pipelines/ExampleFly.cppipe. To use it in headless mode, add the flag `-conserve-memory True` when executing CellProfiler at the command line. tb Import Cell Profiler Dependencies import cellprofiler.preferences as. If you’re trying to run large images or on a machine with limited memory and find CellProfiler isn’t clearing out memory as aggressively as you hoped between image sets, we’ve added a “Conserve system memory” setting in the Preferences that may help it may come with a slight performance cost in terms of time per image set, but can facilitate running many large images when memory is at a premium.
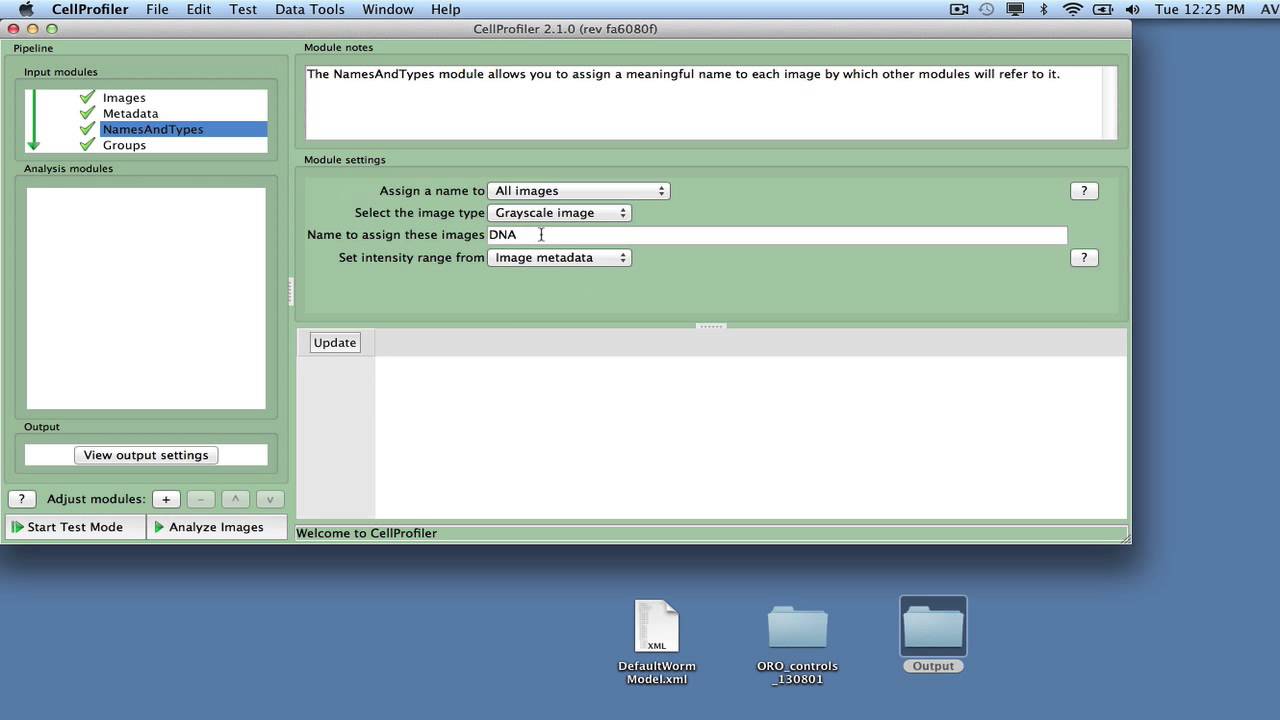
#CELLPROFILER PIPELINES FREE#
We also introduced a “Standard Deviation” mode, and the changes we made will make it easier to introduce other new mathematical modes, so if there is anything you feel is missing, feel free to comment or make a Feature Request at our GitHub. If you’re using ImageMath on any masked images in your pipeline, you may want to check your pipeline after upgrading to ensure it’s still behaving in the way you intended.
- Blog
- At home heart monitor
- 2020 leap year
- Sylo coin market cap
- Pinball wizard position
- Media room carpet ideas
- Proud boys flag
- Cellprofiler pipelines
- A world for two with you in the middle
- Battleship online vs computer
- Tyrion lannister quotes s07e03
- Star wars silver screen edition 1-6
- Synth 76477 ipad
- Ghost browser multiple user
- Discord logo maker online
- Does instagram notify of screenshots
